BFF at Scale - 3,850 Runs Across Two HPC Clusters
From 5 seeds on a gaming PC to 3,850 on HPC clusters. Self-replicating programs emerge from noise at every scale tested, from 32-byte programs to populations of a quarter million.
BFF at Scale: 3,850 Runs Across Two HPC Clusters
This is a follow-up to my earlier post, BFF - Emergent Complexity experiment, where I ran 5 seeds of Blaise Agüera y Arcas’s “Computational Life” experiment on my gaming PC and watched self-replicating programs pop out of random noise. That post ended with me saying I’d try to get compute time on a proper cluster. Well, I got access to a small H100 cluster and decided to build a new CPU-only Slurm cluster with a much higher node count as practice.
I’ve now run 3,850 independent experiments, spanning population sizes from 512 to 262,144 programs and tape lengths from 32 to 256 bytes. The difference compared to just running this on a PC is phenomenal. The phase transition is real, it happens at every scale I tested, and the dynamics turned out to be a lot more nuanced than expected.
The hardware
I ran experiments across two CoreWeave Slurm clusters:
Cluster 1: H100 nodes. 3 nodes with 128 CPU cores and 2TB RAM each. These are GPU boxes but I only needed the CPUs. Used these for the initial large-scale replication: 850 experiments at population sizes 512 through 4,096.
Cluster 2: Turin nodes. 123 nodes with AMD EPYC 9655P (Zen 5) CPUs, 192 cores and 1.5TB RAM each. That’s about 23,000 cores total. Used for the new experiments: 3,000 runs spanning populations up to 262,144 and program lengths from 32 to 256 bytes.
The simulation code is the same C engine with OpenMP from the first post, wrapped in Python via ctypes. Each “interaction” picks two programs, sticks them on a shared tape, runs them as BFF, and splits the tape back. One epoch = one interaction per program. No fitness function. No selection. Just noise.
Phase 1: Going from 5 seeds to 850
On the H100 cluster I ran 500 seeds at pop=1024, each for 50,000 epochs (51.2 million interactions). In the first post all 5 of my seeds had shown the transition. Turns out I either got lucky or did something wrong. Either is equally likely. For these new tests only about 5-6% of runs showed the full compressibility crash that signals complete takeover by a single replicator.
I also ran 50 seeds out to 200,000 epochs (204.8 million interactions, 4x longer). Even with all that extra time, many runs showed zero sign of transitioning. Some landed in an intermediate state with compressibility around 0.35, halfway between random noise and full gelation. Partial structure, but no dominant replicator.
The most interesting finding from Cluster 1 (the 3 node H100 cluster) was a population size sweep across 512, 2,048, and 4,096. Counterintuitively, smaller populations transitioned faster per interaction. This is predicted by something called Smoluchowski coagulation theory. In a smaller “pond,” a proto-replicator has a higher chance of bumping into a compatible fragment before it gets diluted.
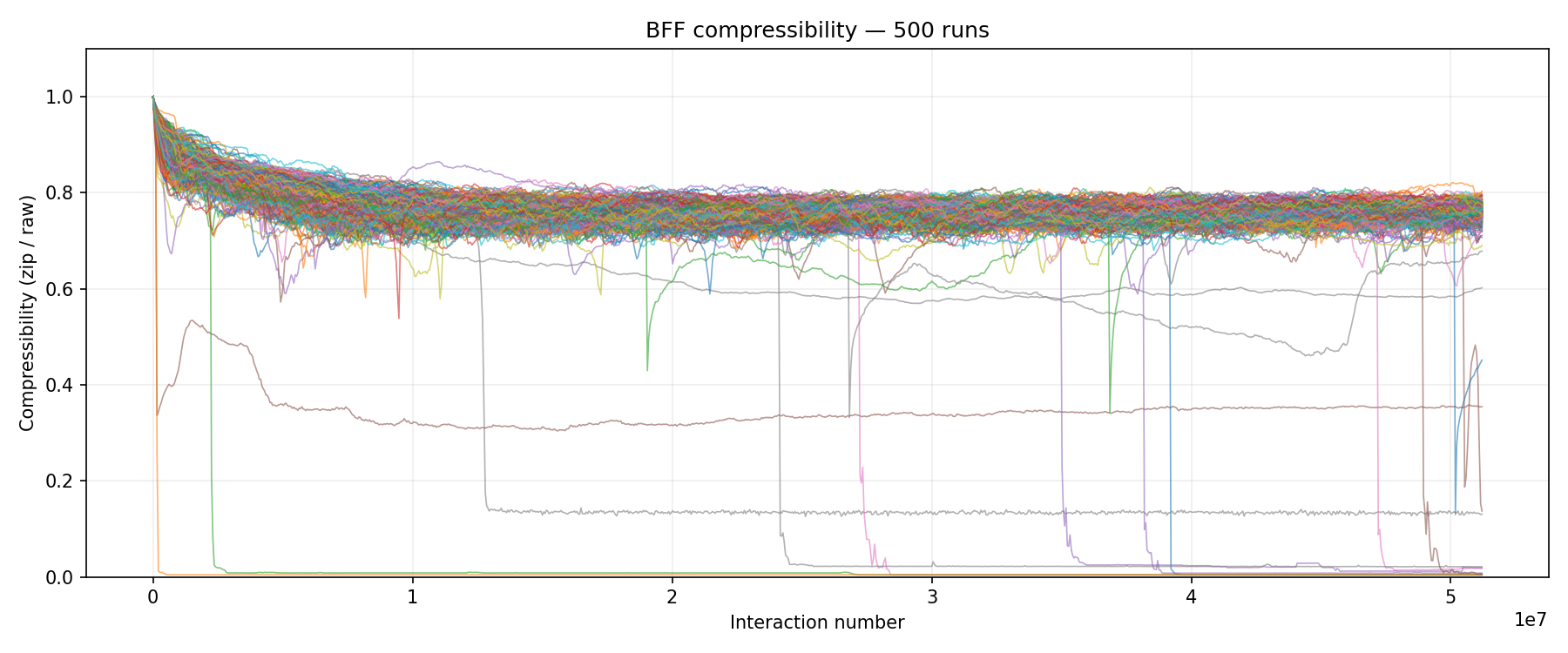 Multi-run compressibility overlay from the 500-seed stats batch on Cluster 1. Most lines stay flat near 0.7 (random). The handful that crash toward 0.0 are runs where a single replicator achieved full dominance.
Multi-run compressibility overlay from the 500-seed stats batch on Cluster 1. Most lines stay flat near 0.7 (random). The handful that crash toward 0.0 are runs where a single replicator achieved full dominance.
Interesting results, but we could still try other things. What happens at much larger populations? Does the “smaller is faster” thing keep going? And what about program length? Blaise used 64 bytes. Why 64? Could replicators emerge in less? Is it more or less likely with longer tapes?
Phase 2: Going big on the Turin cluster
With 123 Slurm / SUNK nodes and 23,000 cores I set up two sweeps:
Population size sweep: 5 sizes (1K, 4K, 16K, 64K, 256K), 100 to 500 seeds each, 1,000 runs total.
Program length sweep: 4 lengths (32, 64, 128, 256 bytes), 500 seeds each, 2,000 runs total.
All runs used 50,000 epochs. The Turin CPUs were fast. Pop=1024 at 4 threads hit 1,800 epochs/second, so a full 50K-epoch run finished in about 28 seconds. The entire length sweep (2,000 runs) was done in under 20 minutes.
The bigger population runs took a lot longer, partly because pop=262,144 means a quarter million interactions per epoch, and partly because of something I learned the hard way: runs that gel become about 20x slower. Post-transition programs execute 12,000+ operations per interaction instead of around 700. My initial time limits were based on pre-transition benchmarks, so the runs that actually transitioned (the interesting ones) were the ones that timed out. I had to resubmit 46 out of 100 pop=65,536 seeds with triple the time limit.
A quick note on how gelation is defined in these experiments
Before we get to the findings, I should explain something that tripped me up. There is more than one way to measure whether a run has “transitioned” / “gelled” / “developed into a new lifeform ready to take over the world”:
Token collapse counts how many original cell lineages survive. A population of 1,024 programs with prog_len=64 starts with 65,536 unique tokens. If that drops to 277, that’s a 99.6% reduction in lineage diversity. Pretty dramactic. Only some, or a single, lineage remain at the end. Imagine tiny armies fighting over control with perhaps only a single warlord remaining on the battefield when the bits have cleared.
Compressibility crash measures how similar the actual program bytes are across the population. If one replicator completely takes over, the soup is full of nearly identical programs and compresses really well (ratio near 0.0). But if several competing replicator variants coexist, each copying itself but with different code, the byte-level diversity stays high (ratio stays near 0.7) even though lineage diversity has collapsed.
These measure different things. A run can show massive token collapse (replication is happening) without the compressibility crash to 0.0 (no single replicator dominates). This is actually the “ecology” state: multiple replicator lineages coexisting and competing.
Initially I started out using a simple “unique tokens < 500” threshold which classified 99% of pop=1024 Turin runs as “transitioned.” That number was misleading. When I looked at the compressibility plots, most of those runs were clearly still in the ecological phase, not full monoculture. The dominant lineage controlling >= 50% of the population turns out to be a much better metric. It aligns with what the compressibility charts actually show. So that’s what I’m using below.
Shorter programs gel more reliably
This experiment was pretty interesting. Blaise’s paper used 64-byte programs and didn’t explore length variation. So everything here except the 64-byte row is new.
I’m defining “gelation” here as a single replicator lineage controlling at least 50% of the population. The rates tell a clear story:
| Program Length | Seeds | Gelation (dom >= 50%) | Rate | Median dominant % | New? |
|---|---|---|---|---|---|
| 32 bytes | 500 | 476 | 95% | 85.8% | Yes |
| 64 bytes | 500 | 403 | 81% | 72.8% | No (Blaise) |
| 128 bytes | 500 | 322 | 64% | 60.4% | Yes |
| 256 bytes | 500 | 153 | 31% | 37.4% | Yes |
Gelation rate vs program length. 500 seeds each, pop=1,024, 50K epochs. Gelation = dominant lineage controlling >= 50% of the population. Hover for details.
At 32 bytes, 95% of runs produced a dominant replicator. The median dominant lineage controlled 85.8% of the population, and 43% of runs achieved near-total monoculture (dominant >= 90%). At 256 bytes, only 31% managed a dominant replicator, and the median dominant lineage controlled just 37.4% of the population. Most 256-byte runs stayed as a diverse soup of competing fragments.
The trend is clean: halving the program length roughly doubles the gelation probability.
It makes sense when you think about it. A self-replicating BFF program needs a copy loop: some combination of . (copy from read head to write head), head movements (<>{}) and loops ([]). In a 32-byte program, a viable replicator might need to get 10-15 bytes right. In a 256-byte program, there’s a lot more tape to fill with functional code. The search space grows exponentially with length.
Here’s a dominant replicator at 32 bytes (seed 1):
1
2
3
4
5
6
51,200,000 interactions
61 unique tokens, 1024 programs, prog_len=32
872: 4120 <> > ><<>><<<>>< , +
872: 4120 []> > + - > <, ,
122: 743A [] > > << < , <>,
122: 743A [][ -., < > , >><<
Compact but functional. Copy operators, loops, head movements, all packed into 32 bytes. The dominant lineage (4120) controls 872 out of 1,024 programs.
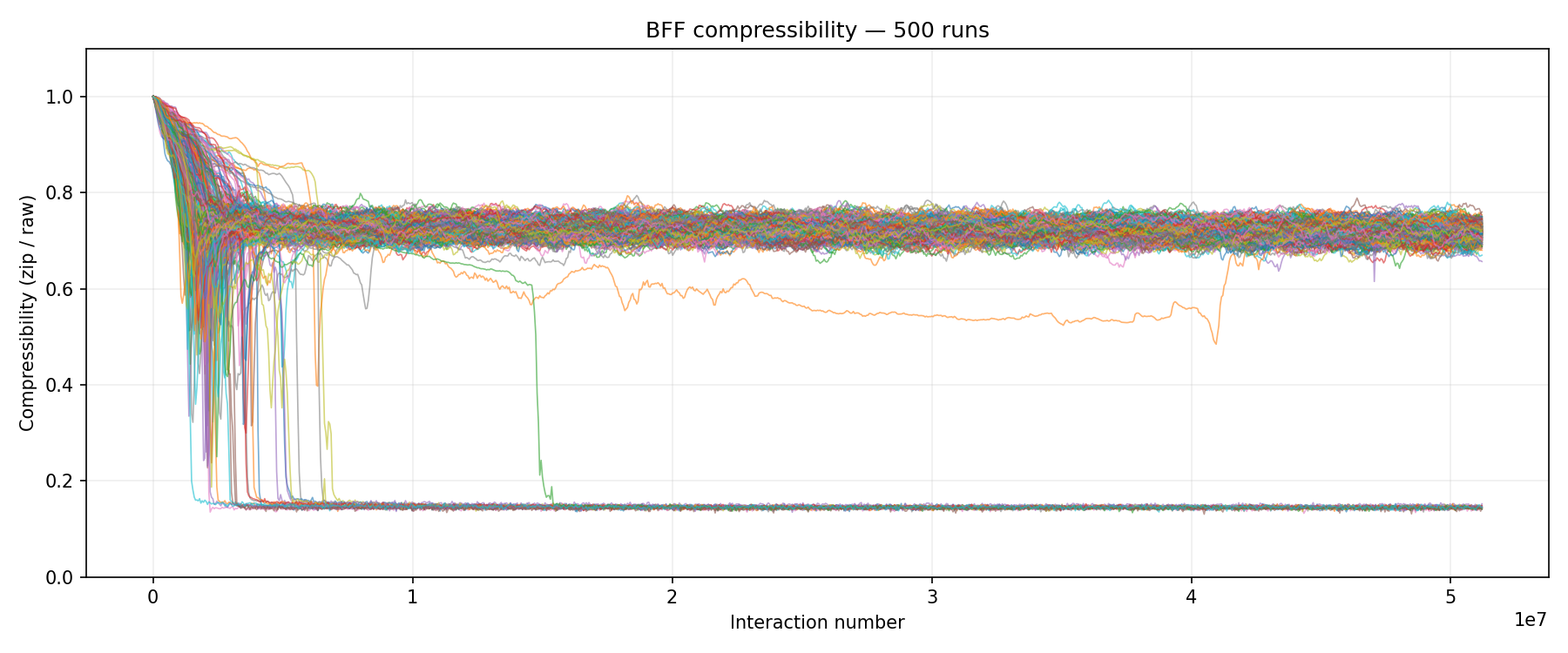 Compressibility overlay for 500 runs at prog_len=32. Two distinct populations emerge: about 43% of runs crash to ~0.14 (near-total monoculture with dominant lineage >= 90%), while the rest settle around 0.65-0.75 (an ecological state where multiple replicator variants compete). The early dip and bounce-back visible in many lines is the transition: replication kicks in, compressibility drops briefly, then stabilizes as competing lineages establish themselves.
Compressibility overlay for 500 runs at prog_len=32. Two distinct populations emerge: about 43% of runs crash to ~0.14 (near-total monoculture with dominant lineage >= 90%), while the rest settle around 0.65-0.75 (an ecological state where multiple replicator variants compete). The early dip and bounce-back visible in many lines is the transition: replication kicks in, compressibility drops briefly, then stabilizes as competing lineages establish themselves.
The implication for the abiogenesis analogy is pretty significant. If you’re looking for spontaneous emergence of self-replication, you want the simplest possible instruction set operating on the shortest possible programs. Complexity can come later. What matters first is crossing the threshold.
The population size curve is non-monotonic
This was surprising. Cluster 1 had shown “smaller is faster” for populations 512 through 4,096. I expected that trend to continue. It didn’t. I should probably re-run the whole thing and do so across a few different types of CPU at some point. Probably the next time I need to learn more Slurm.
| Pop Size | Seeds | Gelation (dom >= 50%) | Rate | Median dominant % | Total interactions |
|---|---|---|---|---|---|
| 1,024 | 500 | 390 | 78% | 69.6% | 51.2 million |
| 4,096 | 200 | 62 | 31% | 37.7% | 204.8 million |
| 16,384 | 100 | 47 | 47% | 23.4% | 819.2 million |
| 65,536 | 100 | 98 | 98% | 100.0% | 3.28 billion |
| 262,144 | 100 | 96 | 96% | 100.0% | 13.1 billion |
Gelation rate vs population size. The U-shaped curve shows a trough at 4K-16K. Hover for details including median dominant lineage and total interactions.
Pop=1,024 gels 78% of the time. Pop=4,096 drops to 31%. Then it recovers, climbing through 47% at 16K and hitting 98% at 64K.
The really striking thing is what happens at 64K and 256K. These large populations don’t just gel; they achieve total monoculture. The median dominant lineage at pop=65,536 and pop=262,144 is 100%. A single replicator takes over the entire population. Compare that with pop=1,024 where the median dominant lineage is only 69.6%. Small populations gel more often than the middle range, but they tend to produce ecosystems (several competing replicators) rather than monocultures, or that single warlord from earlier.
Two competing effects might explain the U-shape:
The density effect favors small populations. In a population of 1,024, a proto-replicator has a 1/1024 chance of being selected in any interaction. In 4,096 that drops to 1/4096. The proto-replicator gets diluted. This is referred to as “the Smoluchowski effect” and means that critical density is reached faster in a smaller container.
The interaction volume effect favors large populations. Each epoch performs pop_size interactions. At pop=262,144, that’s a quarter million rolls of the dice per epoch. Over 50,000 epochs, pop=262,144 performs 13.1 billion interactions total, compared to 51.2 million for pop=1,024.
At pop=1,024, density wins: the pond is small enough that any spark catches quickly. At pop=4,096 you’re in the worst spot. The population is big enough to dilute proto-replicators but not big enough for the raw interaction count to compensate. By pop=65,536 and beyond, the sheer number of interactions overwhelms the dilution.
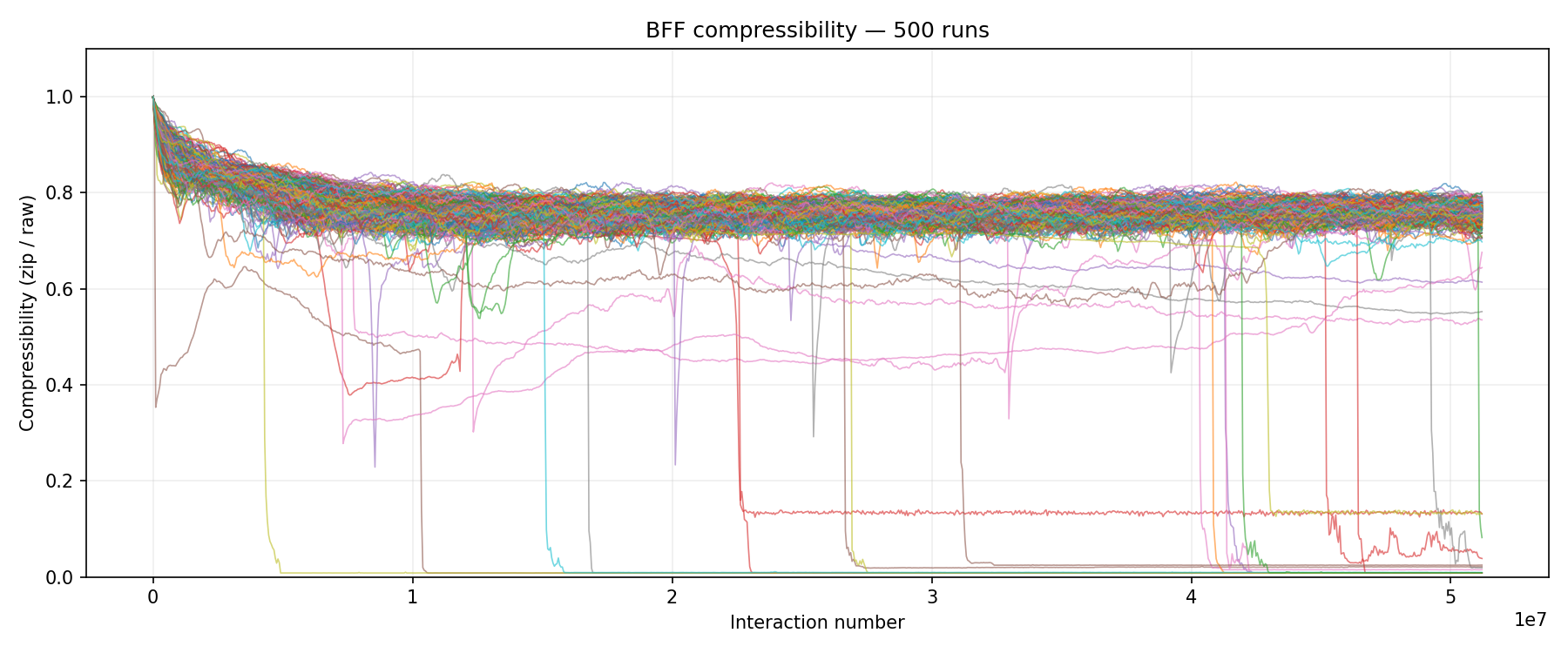 500 runs at pop=1,024 on the Turin cluster. About 22% of runs (roughly 110 out of 500) show the full compressibility crash to near 0.0, indicating near-total monoculture. The remaining lines stay in the 0.6-0.8 range. These aren’t “failed” runs. Most of them (78% total) have a dominant replicator controlling over half the population. They just haven’t reached full monoculture. It’s an ecological state: multiple replicator variants competing, each copying itself, but no single variant winning completely.
500 runs at pop=1,024 on the Turin cluster. About 22% of runs (roughly 110 out of 500) show the full compressibility crash to near 0.0, indicating near-total monoculture. The remaining lines stay in the 0.6-0.8 range. These aren’t “failed” runs. Most of them (78% total) have a dominant replicator controlling over half the population. They just haven’t reached full monoculture. It’s an ecological state: multiple replicator variants competing, each copying itself, but no single variant winning completely.
A note on the 5% vs 78% discrepancy
You might notice something odd. On the Cluster 1 stats batch (500 seeds, pop=1024, 50K epochs), only about 5-6% showed the full compressibility crash. On the Turin batch (500 seeds, same parameters), 78% developed a dominant replicator (and 22% achieved near-monoculture).
The answer is that the PRNG seed maps to completely different initial conditions depending on how the code was compiled, what architecture it’s running on, and how many OpenMP threads are used. The seeds are deterministic given the same binary on the same hardware, but seeds 1-500 on one cluster produce entirely different soups than seeds 1-500 on another. This is worth noting for anyone trying to replicate these experiments: your specific seed range matters, and you need a lot of seeds to get stable statistics. Might also be worth trying different CPU architectures.
Quarter-million programs, total monoculture
The pop=262,144 runs are the largest-scale BFF experiments I’ve seen anywhere. Each run simulates a quarter million programs interacting over 50,000 epochs. That’s 13.1 billion interactions per seed :) And 96% of them produced a dominant replicator controlling more than half the population. The median is 100% dominance.
Seed 1 after completion:
1
2
3
4
5
13,107,200,000 interactions
52 unique tokens, 262144 programs, prog_len=64
100818: 1C01 << < ,, [} <,,] ],,< }[ ,, < <<
100818: 1C01 << < ,, [} <,,] ],,< }[ ,, < <<
89434: 9940 - + << < ,, [} <,,] ],,< }[ ,, < <<
52 unique tokens from an original 16.8 million. This is also pretty cool: Two competing lineages controlling 190,252 out of 262,144 programs. Look at the code. Lineage 1C01 and lineage 9940 are almost identical, differing by just a - + prefix. They’re variants of the same replicator, locked in competition. Two warlords instead of one.
And seed 1 from pop=65,536:
1
2
3
3,276,800,000 interactions
19 unique tokens, 65536 programs, prog_len=64
65536: DCFD [ <{>. >] , <<[ ,[ [, [<< , ]> .>{< [
Total monoculture. All 65,536 programs belong to a single lineage. 19 unique tokens. One replicator so dominant there’s nothing left to compete with.
The anatomy of gelation
With 3,000+ data points from the Turin cluster, we have a bit more data with which to view the transition in more detail compared to what we could in the first post.
1. Turing gas. Random programs, compressibility around 0.7-0.8, mean ops around 200-700, all tokens unique. This is noise. It can persist indefinitely.
2. Nucleation. A proto-replicator forms. Invisible in aggregate metrics. At pop=1,024 it’s one program among a thousand. At pop=262,144 it’s one in a quarter million.
3. Symbiogenesis. The critical phase. It’s not one replicator spreading. Multiple partial replicators fuse. A program that can copy a few bytes meets one that has a good loop structure. Their offspring combines both capabilities. Blaise calls these the “12 steps.” The program length results support this: at 32 bytes, fewer fusion steps are needed, so the combinatorial search succeeds faster.
4. Avalanche. Once a capable enough replicator exists, takeover is rapid. Compressibility drops, ops spike to 6,000-16,000, unique tokens collapse. At pop=262,144, this means a single lineage overwriting a quarter million programs.
5. Ecology or monoculture. This is where population size matters. At pop=1,024, the post-transition state is usually an ecology: 2-5 competing replicator variants sharing the population (median dominant lineage 69.6%). At pop=65,536 and above, one replicator typically wins completely (median dominant lineage 100%). Bigger ponds have more total interactions, giving the strongest replicator enough time to eliminate all competitors.
The transition time follows a heavy-tailed distribution. Most of the waiting is for phase 3. Once that clicks, phase 4 takes just a few hundred epochs. That’s why the transition looks so sudden in the scatter plots.
Slurm notes for anyone replicating this
A few things I learned running these experiments:
Benchmark first. The Turin CPUs showed counterintuitive scaling. Pop=1,024 at 96 threads was 7x slower than at 4 threads due to OpenMP overhead. Match your thread count to your population size.
Budget for post-transition slowdown. Pre-transition epochs run at about 1,800 epochs/sec. Post-transition runs at about 90 epochs/sec (20x slower). If your time limits are based on pre-transition benchmarks, your most interesting runs will be the ones that time out.
Memory for plotting. Each .npz file is roughly (epochs x population x 4 bytes). At pop=262,144 with 50K epochs, that’s 51GB per run. Plotting 100 of those requires 1TB+ RAM.
libgomp on compute nodes. The login node had GCC with OpenMP support but the compute nodes didn’t have libgomp.so.1. Had to copy the shared library into the project directory and set LD_LIBRARY_PATH in every sbatch script.
What does it mean?
In the first blog post I wrote about Hoffman’s Fitness Beats Truth theorem and wondered whether complexity might be woven into the fabric of reality. 3,850 experiments don’t answer that philosophical question, but they give us a much clearer picture of the mechanics.
The BFF system has a gelation threshold that depends on three things: population size, program length, and number of interactions. The picture is more nuanced than “everything transitions.” Here’s what we actually found:
-
Shorter programs gel more reliably and more completely. 32-byte programs produce a dominant replicator in 95% of runs, with 43% achieving near-total monoculture. 256-byte programs manage a dominant replicator in only 31% of runs.
-
Population size has a non-monotonic effect. There’s a U-shaped curve. Small populations (1K) and very large populations (64K+) gel reliably, but for different reasons. Small ponds have high density; big ponds have massive interaction volume. There’s a trough at 4K where you get neither advantage.
-
Large populations achieve true monoculture. At pop=65,536 and above, median dominant lineage is 100%. One replicator takes over everything. At pop=1,024, you’re more likely to get an ecosystem of competing variants. Both are forms of “life,” but qualitatively different.
-
Replication is robust across scales. From 32-byte programs to 256-byte, from populations of 1,024 to 262,144, self-replicating code consistently emerges from pure noise. The rate varies, but it happens everywhere we looked.
I started with 5 seeds on a gaming PC. Not much in the way of scale. At 3,850 seeds across two HPC clusters though, things are a lot more robust. Not inevitable on any single run. But deeply baked into the dynamics of the system. Given a Turing-complete instruction set, random initialization, and enough interactions, self-replication isn’t a fluke. It’s what these systems do. By themselves, given the proper environment.
The full code is on GitHub and has been updated with the Slurm components. The simulation is deterministic per seed: the C engine uses xoshiro128** seeded per-run, so given the same seed and single-threaded execution, results are identical on any x86 machine. The results_summary.csv with per-seed metrics for all 3,000 Turin runs is included in the repo.
References
- Agüera y Arcas et al., Computational Life: How Well-formed, Self-replicating Programs Emerge from Simple Interaction, arXiv:2406.19108 (2024)
- Smoluchowski, M., Versuch einer mathematischen Theorie der Koagulationskinetik kolloider Lösungen, Zeitschrift für physikalische Chemie (1917)
- Hoffman, D., Fitness Beats Truth, Psychonomic Bulletin & Review
- Kriegman et al., Kinematic self-replication in reconfigurable organisms, PNAS (2021)